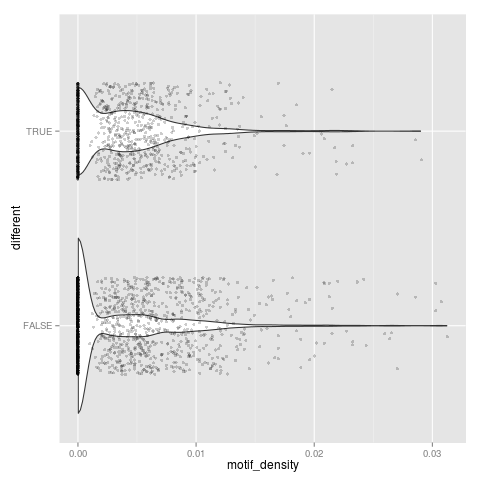

t-test: p=2.2271e-05
Neutral set average density: 0.00281495
Different set average density: 0.00354888
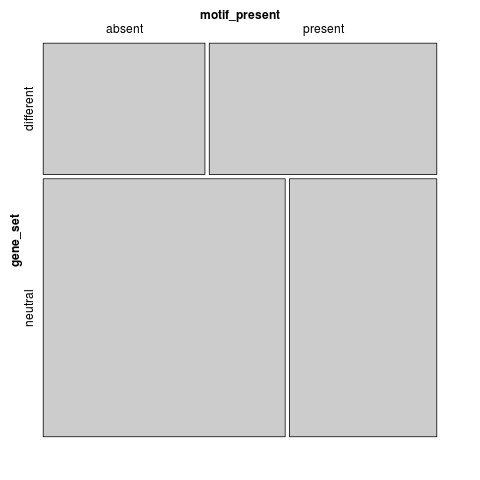
motif_present gene_set absent present different 413 581 neutral 1214 739
Fisher's exact test: p=2.28459e-26
Binomial GLM: p=5.79622e-26
Call:
glm(formula = motif_present ~ different, family = binomial, data = daf)
Deviance Residuals:
Min 1Q Median 3Q Max
-1.3254 -0.9751 -0.9751 1.3942 1.3942
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -0.49638 0.04666 -10.64 <2e-16 ***
differentTRUE 0.83768 0.07949 10.54 <2e-16 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 4053.4 on 2946 degrees of freedom
Residual deviance: 3940.2 on 2945 degrees of freedom
AIC: 3944.2
Number of Fisher Scoring iterations: 4
Binomial GLM adjusted for length: p=0.0322352
Call:
glm(formula = motif_present ~ length + different, family = binomial,
data = daf)
Deviance Residuals:
Min 1Q Median 3Q Max
-3.0452 -0.8236 -0.6340 0.9659 1.8874
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) -1.8508340 0.0801453 -23.093 <2e-16 ***
length 0.0072620 0.0003489 20.817 <2e-16 ***
differentTRUE 0.2004542 0.0936053 2.141 0.0322 *
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 4053.4 on 2946 degrees of freedom
Residual deviance: 3255.6 on 2944 degrees of freedom
AIC: 3261.6
Number of Fisher Scoring iterations: 5